Protein-Protein Interactions
Protein-protein interactions (PPIs) are intrinsic to all cellular processes, driving both metabolic and regulatory pathways. Despite the numerous techniques available, detection of transient short-lived PPIs remains challenging4. The main fluorescence microscopic techniques developed for visualizing PPIs in a cellular context are based on Föster resonance energy transfer (FRET) or bimolecular fluorescence complementation (BiFC)5. Both techniques detect the interaction between a pair of labeled molecules. Although highly informative, they require fine positioning of the labels and in the majority of the applications the spatial resolution achieved is limited by the diffraction of light to about 200 nm. More information concerning the use of FRET to detect PPIs can be found at Cellular Signalling).
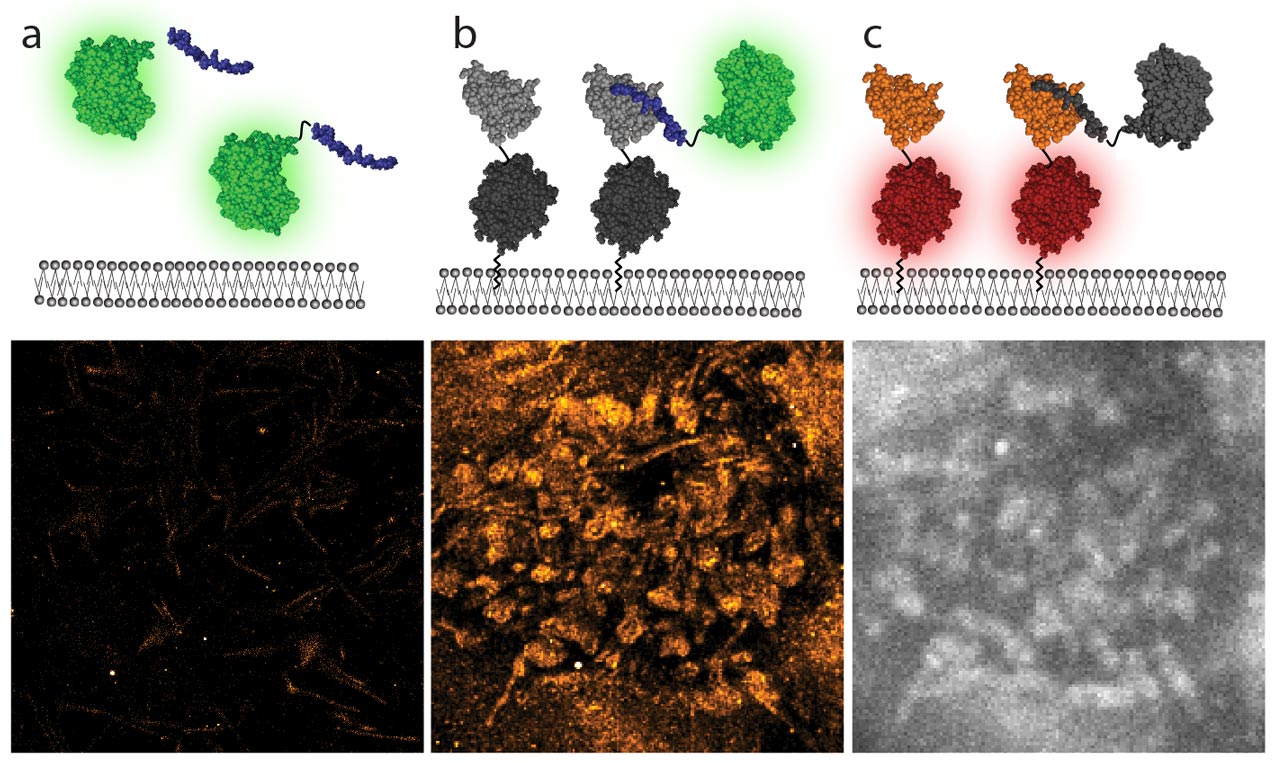
We have use a single molecule localization based super-resolution technique to detect and map PPIs at the cell membrane. This new variant of PAINT that enables mapping of short-lived transient interactions between cytosolic and membrane-bound proteins inside living mammalian cells, at the nanometer scale. In this method the protein of interest is labeled with a light-controllable fluorescent protein and imaged under TIRF illumination, which leads to the selective activation and subsequent detection of molecules in close proximity with the plasma membrane. Interacting molecules are discriminated using a stringent fitting of the fluorescence signal recorded for every single molecule.